Automatic detection of diseases by use of computers is an important, but still unexplored field of research. Such innovations may improve medical practice and refine health care systems all over the world. However, datasets containing medical images are hardly available, making reproducibility and comparison of approaches almost impossible. Here, we present Kvasir, a dataset containing images from inside the gastrointestinal (GI) tract. The collection of images are classified into three important anatomical landmarks and three clinically significant findings. In addition, it contains two categories of images related to endoscopic polyp removal. Sorting and annotation of the dataset is performed by medical doctors (ex- perienced endoscopists). In this respect, Kvasir is important for research on both single- and multi-disease computer aided detec- tion. By providing it, we invite and enable multimedia researcher into the medical domain of detection and retrieval.
The human digestive system may be affected by several diseases. Altogether esophageal, stomach and colorectal cancer accounts for about 2.8 million new cases and 1.8 million deaths per year. Endoscopic examinations are the gold standards for investigation of the GI tract. Gastroscopy is an examination of the upper GI tract including esophagus, stomach and first part of small bowel, while colonoscopy covers the large bowel (colon) and rectum. Both these examinations are real-time video examinations of the inside of the GI tract by use of digital high definition endoscopes. Endoscopic examinations are resource demanding and requires both expensive technical equipment and trained personnel. For colorectal cancer prevention, endoscopic detection and removal of possible precancerous lesions are essential. Adenoma detection is therefore considered to be an important quality indicator in colorectal cancer screening. However, the ability to detect adenomas varies between doctors, and this may eventually affect the individuals’ risk of getting colorectal cancer. Endoscopic assessment of severity and sub-classification of different findings may also vary from one doctor to another. Accurate grading of diseases are important since it may influence decision-making on treatment and follow-up. For example, the degree of inflammation directly affects the choice of therapy in inflammatory bowel diseases (IBD). An objective and automated scoring system would therefore be highly welcomed. Automatic detection, recognition and assessment of pathological findings will probably contribute to reduce inequalities, improve quality and optimize use of scarce medical resources. Furthermore, since endoscopic examinations are real-time investigations, both normal and abnormal findings have to be recorded and documented within written reports. Automatic report generation would proba- bly contribute to reduce doctors’ time required for paperwork and thereby increase time to patient care. Reliable and careful docu- mentation with the use of minimal standard terminology (MST) may also contribute to improved patient follow-up and treatment. To our knowledge, a standardized and automatic reporting system that ensure high quality endoscopy reports does not exist. In order to make the health care system more scalable and cost effective, basic research in the intersection between computer science and medicine must go beyond traditional medical imaging by combining this area with multimedia data analysis and retrieval, artificial intelligence, and distributed processing. Next-generation medical big-data applications are a frontier for innovation, compe- tition and productivity, where there are currently large initiatives both in the EU and the US. In the area of multimedia research, people are starting to see the synergies between multimedia and medical systems. For automatic algorithmic detection of abnormalities in the GI tract, there have been many proposals from various research communities, especially for the topic of polyp detection. Hovever, the results are hard to reproduce due to lack of available medical data, i.e., the work listed above all use different and non-public data sets. Here, we therefore publish Kvasir: A Multi-Class Image Dataset for Computer Aided Gastrointestinal Disease Detection from the Vestre Viken Health Trust (Norway) containing not only polyps, but also two other findings, two classes related to polyp removal and three anatomical landmarks in the GI tract.
Data Collection
The data is collected using endoscopic equipment at Vestre Viken Health Trust (VV) in Norway. The VV consists of 4 hospitals and provides health care to 470.000 people. One of these hospitals (the Bærum Hospital) has a large gastroenterology department from where training data have been collected and will be provided, making the dataset larger in the future. Furthermore, the images are carefully annotated by one or more medical experts from VV and the Cancer Registry of Norway (CRN). The CRN provides new knowledge about cancer through research on cancer. It is part of South-Eastern Norway Regional Health Authority and is organized as an independent institution under Oslo University Hospital Trust. CRN is responsible for the national cancer screening programmes with the goal to prevent cancer death by discovering cancers or pre-cancerous lesions as early as possible.
Dataset Details
The Kvasir dataset consists of images, annotated and verified by medical doctors (experienced endoscopists), including several classes showing anatomical landmarks, phatological findings or endoscopic procedures in the GI tract, i.e., hundreds of images for each class. The number of images is sufficient to be used for different tasks, e.g., image retrieval, machine learning, deep learning and transfer learning, etc. The anatomical landmarks include Z-line, pylorus, cecum, etc., while the pathological finding includes esophagitis, polyps, ulcerative colitis, etc. In addition, we provide several set of images related to removal of lesions, e.g., "dyed and lifted polyp", the "dyed resection margins", etc. The dataset consist of the images with different resolution from 720x576 up to 1920x1072 pixels and organized in a way where they are sorted in separate folders named accordingly to the content. Some of the included classes of images have a green picture in picture illustrating the position and configuration of the endoscope inside the bowel, by use of an electromagnetic imaging system (ScopeGuide, Olympus Europe) that may support the interpretation of the image. This type of information may be important for later investigations (thus included), but must be handled with care for the detection of the endoscopic findings.
Anatomical Landmarks
An anatomical landmark is a recognizable feature within the GI tract that is easily visible through the endoscope. They are essential for navigating and as a reference point to describe the location of a given finding. The landmarks may also be typical sites for pathology like ulcers or inflammation. A complete endoscopic rapport should preferably contain both brief descriptions and image documentation of the most important anatomical landmarks.
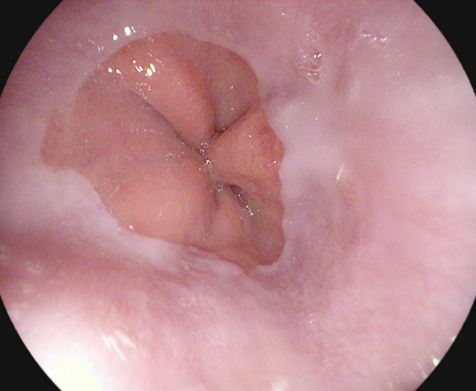 |
Z-line |
The Z-line marks the transition site between the esophagus and the stomach. Endoscopically, it is visible as a clear border where the white mucosa in the esophagus meets the red gastric mucosa. An example of the Z-line is shown in the image to the left. Recognition and assessment of the Z-line is important in order to determine whether disease is present or not. For example, this is the area where signs of gastro-esophageal reflux may appear. The Z-line is also useful as a reference point when describing pathology in the esophagus. |
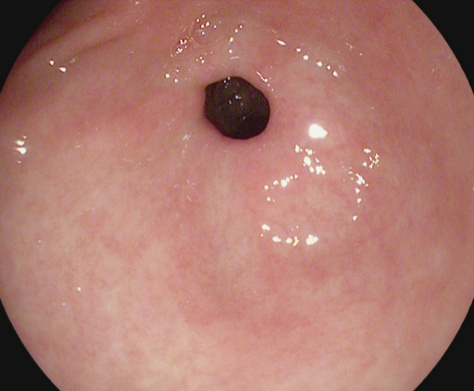 |
Pylorus |
The pylorus is defined as the area around the opening from the stomach into the first part of the small bowel (duodenum). The opening contains circumferential muscles that regulates the movement of food from the stomach. The identification of pylorus is necessary for endoscopic instrumentation to the duodenum, one of the challenging maneuvers within gastroscopy. A complete gastroscopy includes inspection on both sides of the pyloric opening to reveal findings like ulcerations, erosions or stenosis. The image to the left shows an endoscopic image of a normal pylorus viewed from inside the stomach. Here, the smooth, round opening is visible as a dark circle surrounded by homogeneous pink stomach mucosa. |
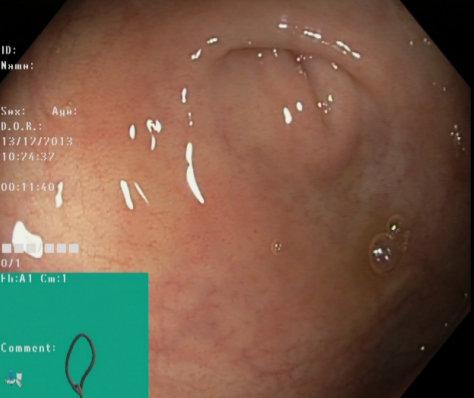 |
Cecum |
The cecum the most proximal part of the large bowel. Reaching cecum is the proof for a complete colonoscopy and completion rate has shown to be a valid quality indicator for colonoscopy. Therefore, recognition and documentation of the cecum is important. One of the characteristics hallmarks of cecum is the appendiceal orifice. This combined with a typical configuration on the electromagnetic scope tracking system may be used as proof for cecum intubation when named or photo documented in the reports. The image to the left shows an example of the appendiceal orifice visible as a crescent shaped slit, and the green picture in picture shows the scope configuration for cecal position. |
Phatological Findings
A pathological finding in this context is an abnormal feature within the gastrointestinal tract. Endoscopically, it is visible as a damage or change in the normal mucosa. The finding may be signs of an ongoing disease or a precursor to for example cancer. Detection and classification of pathology is important in order to initiate correct treatment and/or follow-up of the patient.
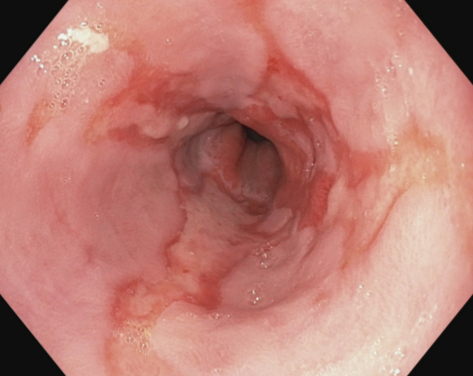 |
Esophagitis |
Esophagitis is an inflammation of the esophagus visible as a break in the esophageal mucosa in relation to the Z-line. The image to the left shows an example with red mucosal tongues projecting up in the white esophageal lining. The grade of inflammation is defined by length of the mucosal breaks and proportion of the circumference involved. This is most commonly caused by conditions where gastric acid flows back into the esophagus as gastroesophageal reflux, vomiting or hernia. Clinically, detection is necessary for treatment initiation to relieve symptoms and prevent further development of possible complications. Computer detection would be of special value in assessing severity and for automatic reporting. |
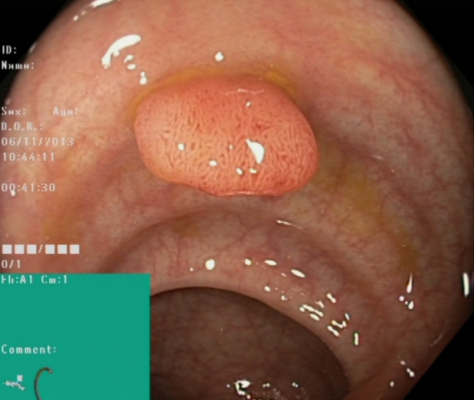 |
Polyps |
Polyps are lesions within the bowel detectable as mucosal outgrows. An example of a typical polyp is shown in the image to the left. The polyps are either flat, elevated or pedunculated, and can be distinguished from normal mucosa by color and surface pattern. Most bowel polyps are harmless, but some have the potential to grow into cancer. Detection and removal of polyps are therefore important to prevent development of colorectal cancer. Since polyps may be overlooked by the doctors, automatic detection would most likely improve examination quality. The green boxes within the image shows an illustration of the endoscope configuration. In live endoscopy, this helps to determine the current localisation of the endoscope-tip (and thereby also the polyp site) within the length of the bowel. Automatic computer aided detection of polyps would be valuable both for diagnosis, assessment and reporting. |
 |
Ulcerative Colitis |
Ulcerative colitis is a chronic inflammatory disease affecting the large bowel. The disease may have a large impact on quality of life, and diagnosis is mainly based on colonoscopic findings. The degree of inflammation varies from none, mild, moderate and severe, all with different endoscopic aspects. For example, in a mild disease, the mucosa appears swollen and red, while in moderate cases, ulcerations are prominent. The image to the left shows an example of ulcerative colitis with bleeding, swelling and ulceration of the mucosa. The white coating visible in the illustration is fibrin covering the wounds. As mentioned earlier, an automatic computer aided assessment system will contribute to more accurate grading of the disease severity. |
Polyp Removal
Polyps in the large bowel may be precursors of cancer and are therefore removed during endoscopy if possible. One of the polyp removal techniques is called endoscopic mucosal resection (EMR). This includes injection of a liquid underneath the polyp, lifting the polyp from the underlying tissue. The polyp is then captured and removed by use of a snare. The lifting minimizes risk of mechanical or electrocautery damage to the deeper layers of the GI wall. Staining dye (i.e., diluted indigo carmine) is added to facilitate accurate identification of the polyp margins. Computer detection of dyed polyps and the site of resection would be important in order to generate computer aided reporting systems for the future.
 |
Dyed and Lifted Polyps |
Figure 9 shows an example of a polyp lifted by injection of saline and indigocarmine. The light blue polyp margins are clearly visible against the darker normal mucosa. Additional valuable information related to automatic reporting may involve successfulness of the lifting and eventual presence of nonlifted areas that might indicate malignancy. |
 |
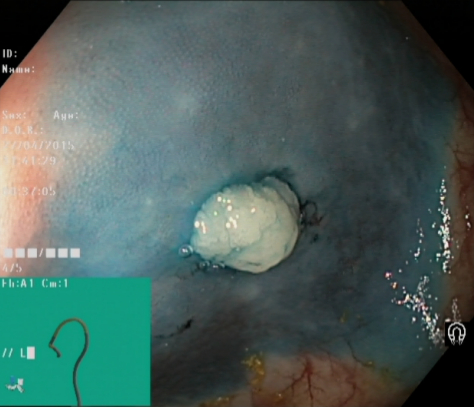
Dyed Resection Margins |
The resection margins are important in order to evaluate whether the polyp is completely removed or not. Residual polyp tissue may lead to continued growth and in worst case malignancy development. Figure 10 illustrates the resection site after removal of a polyp. Automatic recognition of the site of polyp removals are of value for automatic reporting systems and for computer aided assessment on completeness of the polyp removal. |
Applications of the Dataset
Our vision is that the available data may eventually help researchers to develop systems that improve the health-care system in the context of disease detection in videos of the GI tract. Such a system may automate video analysis and endoscopic findings detection in the esophagus, stomach, bowel and rectum. Important results will include higher detection accuracies, reduced manual labor for medical personnel, reduced average cost, less patient discomfort and possibly increased willingness to undertake the examination. In the end, the improved screening will probably significantly reduce mortality and number of luminal GI disease incidents. With respect to direct use in the multimedia research areas, the main application area of Kvasir is automatic detection, classification and localization of endoscopic pathological findings in an image captured in the GI tract. Thus, the provided dataset can be used in several scenarios where the aim is to develop and evaluate algoritmic analysis of images. Using the same collection of data, researchers can easier compare approaches and experimental results, and results can easier be reproduced. In particular, in the area of image retrieval and object detection, Kvasir will play an important initial role where the image collection can be divided into training and test sets for developments of and experiments for various image retrieval and object localization methods including search-based systems, neural-networks, video analysis, information retrieval, machine learning, object detection, deep learning, computer vision, data fusion and big data processing.
Suggested Metrics
Looking at the list of related work in this area, there are a lot of different metrics used, with potentially different names when used in the medical area and the computer science (information retrieval) area. Here, we provide a small list of the most important metrics. For future research, in addition to describing the dataset with respect to total number of images, total number of images in each class and total number of positives, it might be good to provide as many of the metrics below as possible in order to enable an indirect comparison with older work:
- True positive (TP) The number of correctly identified samples. The number of frames with an endoscopic finding which correctly is identified as a frame with an endoscopic finding.
- True negative (TN) The number of correctly identified negative samples, i.e., frames without an endoscopic finding which correctly is identified as a frame without an endoscopic finding.
- False positive (FP) The number of wrongly identified samples, i.e., a commonly called a "false alarm". Frames without an endoscopic finding which is erroneously identified as a frame with an endoscopic finding.
- False negative (FN) The number of wrongly identified negative samples. Frames without an endoscopic finding which erroneously is identified as a frame with an endoscopic finding.
- Recall (REC) This metric is also frequently called sensitivity, probability of detection and true positive rate, and it is the ratio of samples that are correctly identified as positive among all existing positive samples.
- Precision (PREC) This metric is also frequently called the positive predictive value, and shows the ratio of samples that are correctly identified as positive among the returned samples (the fraction of retrieved samples that are relevant).
- Specificity (SPEC) This metric is frequently called the true negative rate, and shows the ratio of negatives that are correctly identified as such (e.g., the fraction of frames without an endoscopic finding are correctly identified as a negative result).
- Accuracy (ACC) The percentage of correctly identified true and false samples.
- Matthews correlation coefficient (MCC) MCC takes into account true and false positives and negatives, and is a balanced measure even if the classes are of very different sizes.
- F1 score (F1) A measure of a test’s accuracy by calculating the harmonic mean of the precision and recall.
In addition to the above metrics, system performance metrics processing speed and resource consumption are of interest. In our work, we have used the achieved frame-rate (FPS) as a metric as real-time feedback is important.
Download Kvasir version 1
| File | Description | Size | Download |
|---|---|---|---|
| kvasir-v1.zip | The entire HyperKvasir dataset in one zip file. | 1.2GB | |
| kvasir-v1-features.zip | The entire HyperKvasir dataset in one zip file. | 4.7MB |
Download Kvasir version 2
| File | Description | Size | Download |
|---|---|---|---|
| kvasir-v2.zip | The entire HyperKvasir dataset in one zip file. | 2.3GB | |
| kvasir-v2-features.zip | The entire HyperKvasir dataset in one zip file. | 9.3MB |
Cite
@inproceedings{Pogorelov:2017:KMI:3083187.3083212,
title = {KVASIR: A Multi-Class Image Dataset for Computer Aided Gastrointestinal Disease Detection},
author = {
Pogorelov, Konstantin and Randel, Kristin Ranheim and Griwodz, Carsten and
Eskeland, Sigrun Losada and de Lange, Thomas and Johansen, Dag and
Spampinato, Concetto and Dang-Nguyen, Duc-Tien and Lux, Mathias and
Schmidt, Peter Thelin and Riegler, Michael and Halvorsen, P{\aa}l
},
booktitle = {Proceedings of the 8th ACM on Multimedia Systems Conference},
series = {MMSys'17},
year = {2017},
isbn = {978-1-4503-5002-0},
location = {Taipei, Taiwan},
pages = {164--169},
numpages = {6},
doi = {10.1145/3083187.3083212},
acmid = {3083212},
publisher = {ACM},
address = {New York, NY, USA},
}
Terms of use
The use of the Kvasir dataset is restricted for research and educational purposes only. The use of the Kvasir dataset for other purposes including commercial purposes is forbidden without prior written permission. In all documents and papers that use or refer to the Kvasir dataset or report experimental results based on the Kvasir dataset, a reference to the dataset paper have to be included.
Contact
Email michael/paalh (at) simula (dot) no if you have any questions about the dataset and our research activities. We always welcome collaboration and joint research!